
Creating publication-ready labels for your outputs
Source:vignettes/articles/improving_readability.qmd
Overview
By default, plot_or() and table_or() display variable names exactly as they appear in your model. Variable names like age_group_35_44 or smoking_status_yes are technical and not suitable for publication-ready outputs.
The solution is to attach human-readable labels to variables. When you do, plotor automatically uses these labels in plots and tables instead of the raw variable names. This approach is much cleaner than manually editing outputs after generation.
Example 1: Oesophageal Cancer Study
The dataset
This case-control study examined oesophageal cancer in Ile-et-Vilaine, France. It contains:
| Variable | Description |
|---|---|
Group |
Case (cancer) or Control (disease-free) |
agegp |
Age group of participant |
alcgp |
Alcohol consumption (grams per day) |
tobgp |
Tobacco consumption (grams per day) |
Preparing the data
# prepare the dataset for modelling
df <-
datasets::esoph |>
# convert aggregated data to tidy observational data
tidyr::pivot_longer(
cols = c(ncases, ncontrols),
names_to = 'Group',
values_to = 'people'
) |>
tidyr::uncount(weights = people) |>
# prepare the variables
dplyr::mutate(
# convert the intervention group to a factor
Group = Group |>
dplyr::recode_values(
"ncases" ~ "Case",
"ncontrols" ~ "Control"
) |>
factor(levels = c("Control", "Case")),
# remove ordering from these predictors
agegp = agegp |> factor(ordered = FALSE),
alcgp = alcgp |> factor(ordered = FALSE),
tobgp = tobgp |> factor(ordered = FALSE)
)
# preview the data
df |> dplyr::glimpse()
#> Rows: 975
#> Columns: 4
#> $ agegp <fct> 25-34, 25-34, 25-34, 25-34, 25-34, 25-34, 25-34, 25-34, 25-34, 2…
#> $ alcgp <fct> 0-39g/day, 0-39g/day, 0-39g/day, 0-39g/day, 0-39g/day, 0-39g/day…
#> $ tobgp <fct> 0-9g/day, 0-9g/day, 0-9g/day, 0-9g/day, 0-9g/day, 0-9g/day, 0-9g…
#> $ Group <fct> Control, Control, Control, Control, Control, Control, Control, C…Without labels
m <- glm(
data = df,
family = "binomial",
formula = Group ~ agegp + alcgp + tobgp
)
# plot the odds ratio with a customised title
plot_or(m)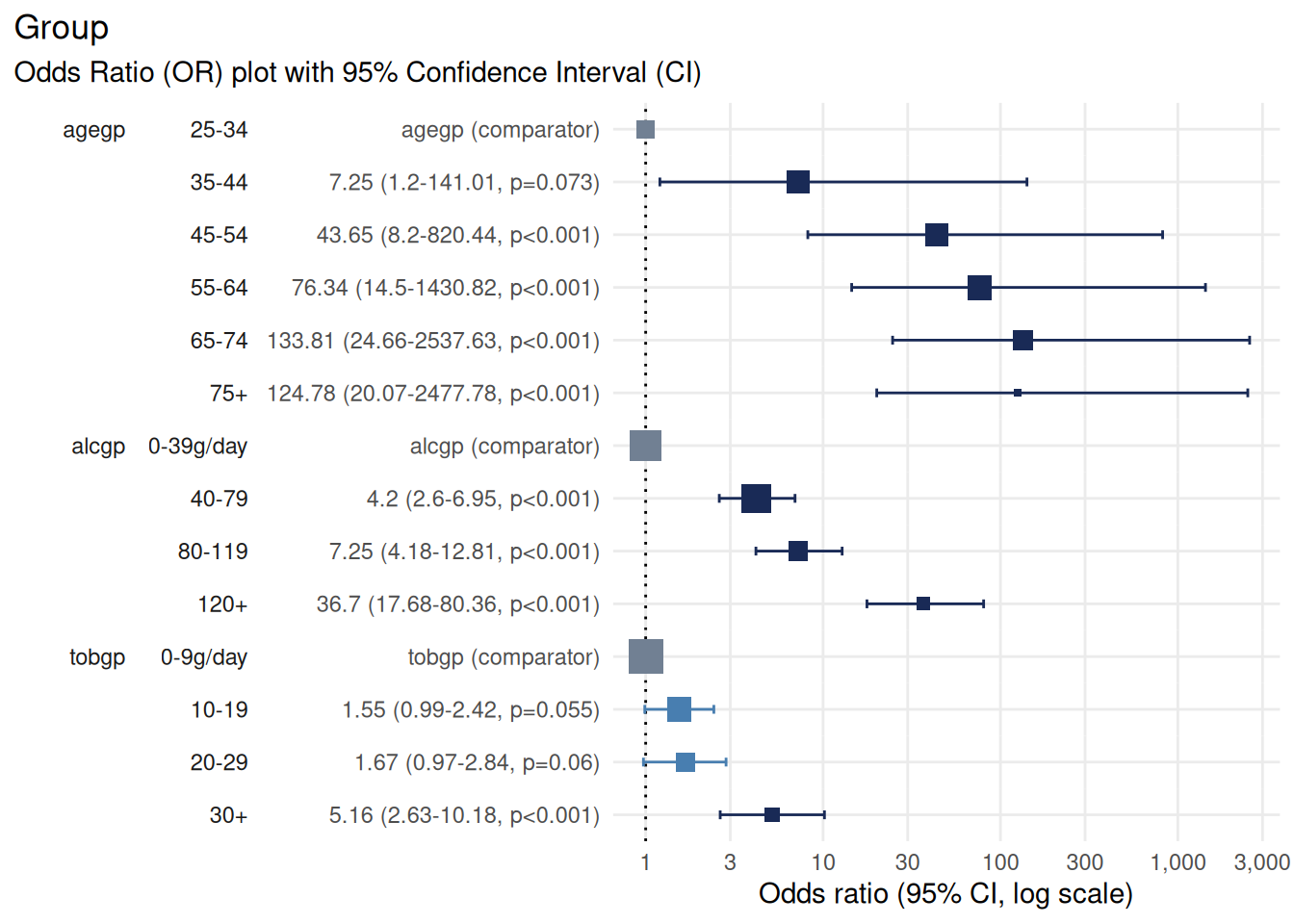
Notice how the plot uses technical variable names like alcgp and tobgp which are not immediately clear to readers.
Adding labels with {labelled}
To make your outputs more readable, attach descriptive labels to your variables before modelling.
First, ensure the package is installed.
install.packages("labelled")
library(labelled)
# create a list that matches variables with user-friendly labels
var_labels <- list(
agegp = "Age group",
alcgp = "Alcohol consumption",
tobgp = "Tobacco consumption",
Group = "Likelihood of developing oesophageal cancer"
)
# apply these variables to our data
labelled::var_label(df) <- var_labels
# preview the data with labels applied
labelled::look_for(df)
#> pos variable label col_type missing
#> 1 agegp Age group fct 0
#>
#>
#>
#>
#>
#> 2 alcgp Alcohol consumption fct 0
#>
#>
#>
#> 3 tobgp Tobacco consumption fct 0
#>
#>
#>
#> 4 Group Likelihood of developing oesophageal ca~ fct 0
#>
#> values
#> 25-34
#> 35-44
#> 45-54
#> 55-64
#> 65-74
#> 75+
#> 0-39g/day
#> 40-79
#> 80-119
#> 120+
#> 0-9g/day
#> 10-19
#> 20-29
#> 30+
#> Control
#> CaseForest plots with labels
Now fit the model using the labelled data:
m <- glm(
data = df,
family = "binomial",
formula = Group ~ agegp + alcgp + tobgp
)
# plot the odds ratio with a customised title
plot_or(m)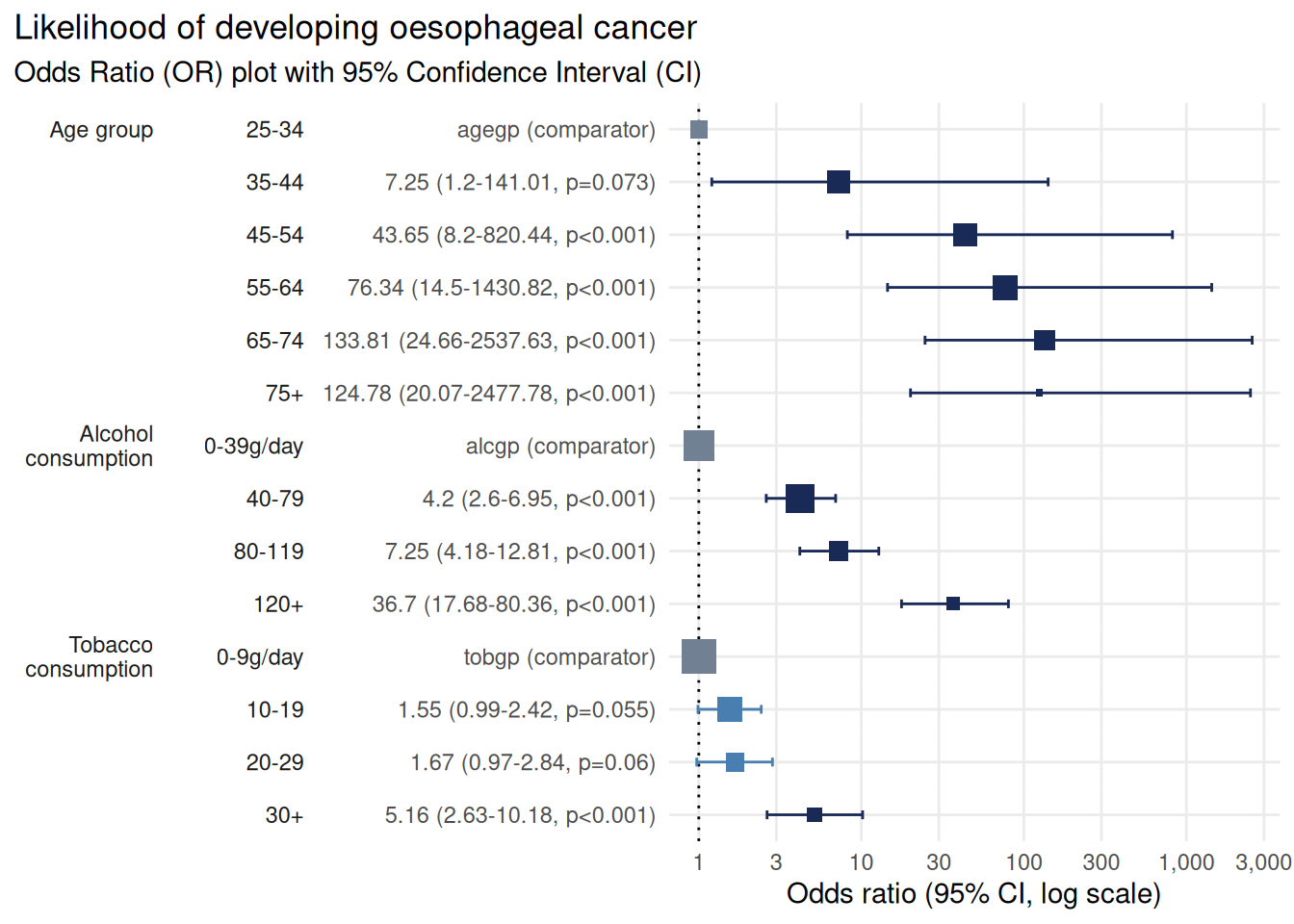
The plot is now much more reader-friendly:
The outcome label (“Likelihood of developing oesophageal cancer”) appears in the title
Predictor labels (“Age group”, “Alcohol consumption”, “Tobacco consumption”) replace technical variable names
Data tables with labels
table_or() also respects variable labels:
table_or(m, output = "gt", assumption_checks = FALSE)| Likelihood of developing oesophageal cancer | ||||||||||||
| Odds Ratio summary table with 95% Confidence Interval | ||||||||||||
Characteristic1
|
Odds Ratio (OR)2
|
95% Confidence Interval (CI)3
|
OR Plot | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Level | N | n | Rate | Class | OR | SE | p | Lower | Upper | Significance | ||
| Age group | 25-34 | 116 | 1 | 0.86% | factor | — | — | — | — | — | Comparator | |
| 35-44 | 199 | 9 | 4.52% | factor | 7.249 | 1.104 | 7.27 × 10−2 | 1.202 | 141.0 | Significant | ||
| 45-54 | 213 | 46 | 21.6% | factor | 43.65 | 1.068 | 4.06 × 10−4 | 8.204 | 820.4 | Significant | ||
| 55-64 | 242 | 76 | 31.4% | factor | 76.34 | 1.065 | 4.68 × 10−5 | 14.50 | 1,431 | Significant | ||
| 65-74 | 161 | 55 | 34.16% | factor | 133.8 | 1.076 | 5.38 × 10−6 | 24.66 | 2,538 | Significant | ||
| 75+ | 44 | 13 | 29.55% | factor | 124.8 | 1.121 | 1.67 × 10−5 | 20.07 | 2,478 | Significant | ||
| Alcohol consumption | 0-39g/day | 415 | 29 | 6.99% | factor | — | — | — | — | — | Comparator | |
| 40-79 | 355 | 75 | 21.13% | factor | 4.198 | 0.2501 | 9.63 × 10−9 | 2.600 | 6.948 | Significant | ||
| 80-119 | 138 | 51 | 36.96% | factor | 7.248 | 0.2848 | 3.51 × 10−12 | 4.183 | 12.81 | Significant | ||
| 120+ | 67 | 45 | 67.16% | factor | 36.70 | 0.3850 | 8.19 × 10−21 | 17.68 | 80.36 | Significant | ||
| Tobacco consumption | 0-9g/day | 525 | 78 | 14.86% | factor | — | — | — | — | — | Comparator | |
| 10-19 | 236 | 58 | 24.58% | factor | 1.550 | 0.2283 | 5.50 × 10−2 | 0.9885 | 2.423 | Not significant | ||
| 20-29 | 132 | 33 | 25% | factor | 1.670 | 0.2730 | 6.04 × 10−2 | 0.9714 | 2.839 | Not significant | ||
| 30+ | 82 | 31 | 37.8% | factor | 5.160 | 0.3441 | 1.85 × 10−6 | 2.631 | 10.18 | Significant | ||
|
1
Characteristics are the explanatory variables in the logistic regression analysis. For categorical variables the first characteristic is designated as a reference against which the others are compared. For numeric variables the results indicate a change per single unit increase.
Level - the name or the description of the explanatory variable. N - the number of observations examined. n - the number of observations resulting in the outcome of interest. Rate - the proportion of observations resulting in the outcome of interest (n / N). Class - description of the data type. |
||||||||||||
|
2
Odds Ratios estimate the relative odds of an outcome with reference to the Characteristic. For categorical data the first level is the reference against which the odds of other levels are compared. Numerical characteristics indicate the change in OR for each additional increase of one unit in the variable.
OR - The Odds Ratio point estimate - values below 1 indicate an inverse relationship whereas values above 1 indicate a positive relationship. Values shown to 4 significant figures. SE - Standard Error of the point estimate. Values shown to 4 significant figures. p - The p-value estimate based on the residual Chi-squared statistic. |
||||||||||||
|
3
Confidence Interval - the range of values likely to contain the OR in 95% of cases if this study were to be repeated multiple times. If the CI touches or crosses the value 1 then it is unlikely the Characteristic is significantly associated with the outcome.
Lower & Upper - The range of values comprising the CI, shown to 4 significant figures. Significance - The statistical significance indicated by the CI, Significant where the CI does not touch or cross the value 1. |
||||||||||||
Example 2: Post-endoscopic pancreatitis study
The dataset
The indo_rct dataset contains details from 602 patients in a randomised controlled study examining indomethacin vs placebo for preventing post-endoscopic pancreatitis.
# prepare the dataset for modelling
df <- medicaldata::indo_rct |>
tibble::as_tibble() |>
# clean up factor levels
dplyr::mutate(
rx = forcats::fct_recode(
.f = rx,
Placebo = "0_placebo", Indomethacin = "1_indomethacin"),
pep = forcats::fct_recode(
.f = pep,
No = "0_no", Yes = "1_yes"
),
amp = forcats::fct_recode(
.f = amp,
No = "0_no", Yes = "1_yes"
)
)
# fit the model without labels
m <- stats::glm(
formula = outcome ~ rx + pep + amp,
family = "binomial",
data = df
)
plot_or(m, assumption_checks = FALSE)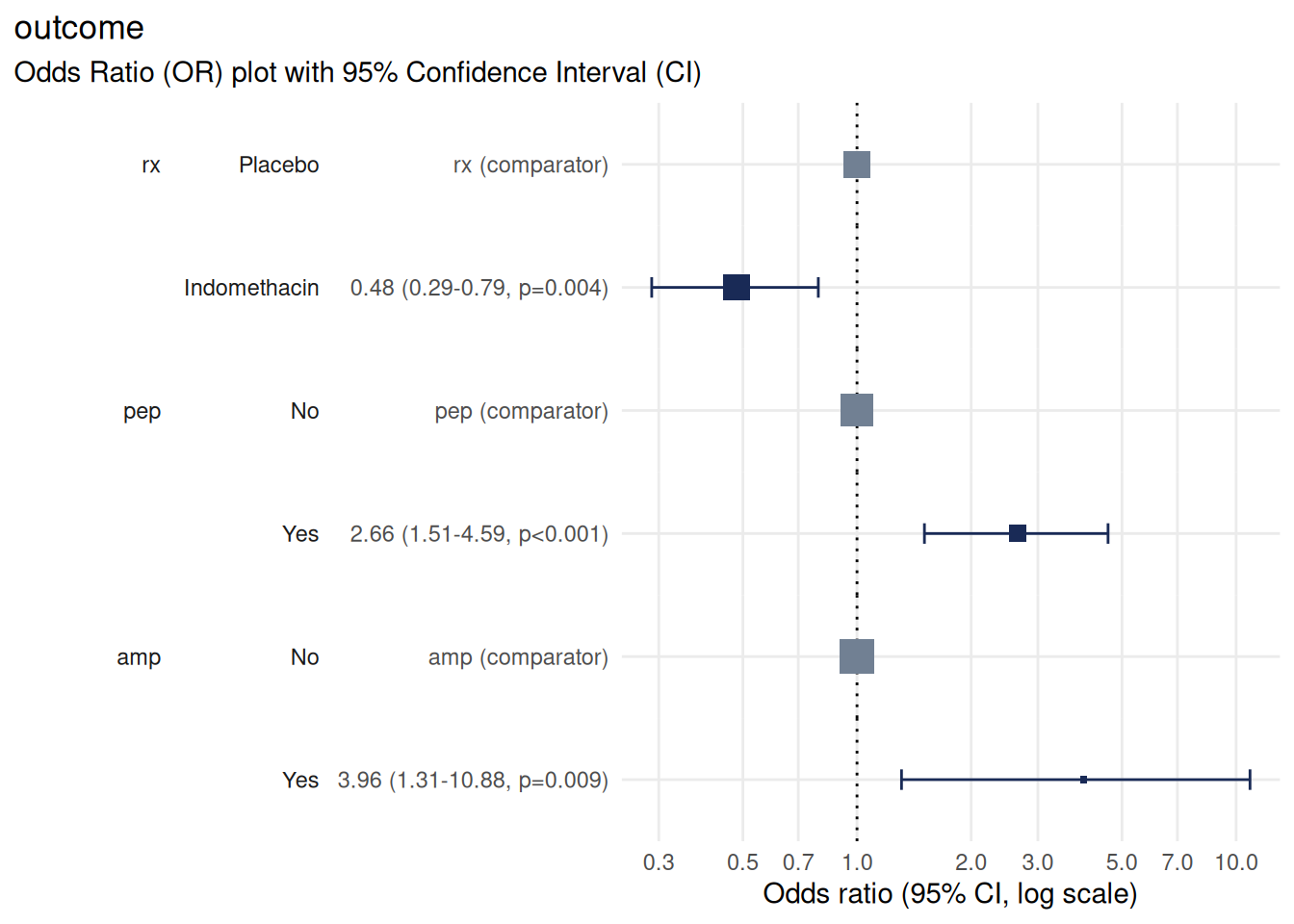
Using {Hmisc} for labels
An alternative to {labelled} is the {Hmisc} package, which is particularly useful if you’re already using other {Hmisc} functions like describe() for summary statistics.
Install it with:
install.packages("Hmisc")
library(Hmisc)Attach labels using the label() function:
# label the variables
Hmisc::label(df$outcome) <- "Likelihood of post-ERCP pancreatitis"
Hmisc::label(df$rx) <- "Treatment arm"
Hmisc::label(df$pep) <- "Previous post-ERCP pancreatitis (PEP)"
Hmisc::label(df$amp) <- "Ampullectomy performed"
# fit the model with labelled data
m <- stats::glm(
formula = outcome ~ rx + pep + amp,
family = "binomial",
data = df
)
# plot
plot_or(m, assumption_checks = FALSE)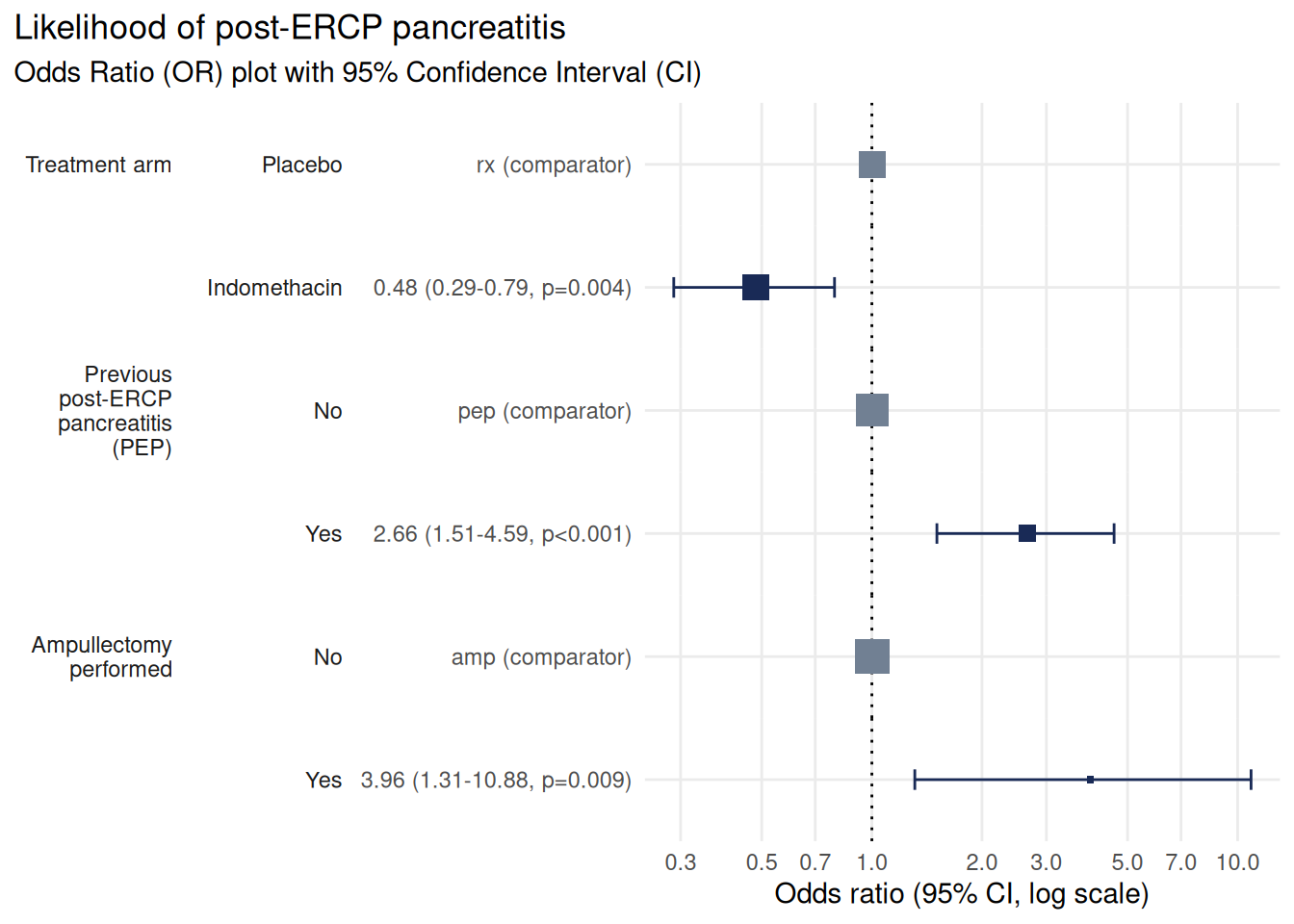
This plot now clearly shows that treatment with Indomethacin has a protective effect against pancreatitis, whereas a history of pancreatitis and ampullectomy are both associated with increased risk.
The same labels also appear in covariate tables:
table_or(m, output = "gt", assumption_checks = FALSE)| Likelihood of post-ERCP pancreatitis | ||||||||||||
| Odds Ratio summary table with 95% Confidence Interval | ||||||||||||
Characteristic1
|
Odds Ratio (OR)2
|
95% Confidence Interval (CI)3
|
OR Plot | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Level | N | n | Rate | Class | OR | SE | p | Lower | Upper | Significance | ||
| Previous post-ERCP pancreatitis (PEP) | No | 506 | 56 | 11.07% | labelled factor | — | — | — | — | — | Comparator | |
| Yes | 96 | 23 | 23.96% | labelled factor | 2.658 | 0.2834 | 5.61 × 10−4 | 1.506 | 4.594 | Significant | ||
| Ampullectomy performed | No | 584 | 73 | 12.5% | labelled factor | — | — | — | — | — | Comparator | |
| Yes | 18 | 6 | 33.33% | labelled factor | 3.960 | 0.5306 | 9.49 × 10−3 | 1.310 | 10.88 | Significant | ||
| Treatment arm | Placebo | 307 | 52 | 16.94% | labelled factor | — | — | — | — | — | Comparator | |
| Indomethacin | 295 | 27 | 9.15% | labelled factor | 0.4816 | 0.2572 | 4.50 × 10−3 | 0.2874 | 0.7905 | Significant | ||
|
1
Characteristics are the explanatory variables in the logistic regression analysis. For categorical variables the first characteristic is designated as a reference against which the others are compared. For numeric variables the results indicate a change per single unit increase.
Level - the name or the description of the explanatory variable. N - the number of observations examined. n - the number of observations resulting in the outcome of interest. Rate - the proportion of observations resulting in the outcome of interest (n / N). Class - description of the data type. |
||||||||||||
|
2
Odds Ratios estimate the relative odds of an outcome with reference to the Characteristic. For categorical data the first level is the reference against which the odds of other levels are compared. Numerical characteristics indicate the change in OR for each additional increase of one unit in the variable.
OR - The Odds Ratio point estimate - values below 1 indicate an inverse relationship whereas values above 1 indicate a positive relationship. Values shown to 4 significant figures. SE - Standard Error of the point estimate. Values shown to 4 significant figures. p - The p-value estimate based on the residual Chi-squared statistic. |
||||||||||||
|
3
Confidence Interval - the range of values likely to contain the OR in 95% of cases if this study were to be repeated multiple times. If the CI touches or crosses the value 1 then it is unlikely the Characteristic is significantly associated with the outcome.
Lower & Upper - The range of values comprising the CI, shown to 4 significant figures. Significance - The statistical significance indicated by the CI, Significant where the CI does not touch or cross the value 1. |
||||||||||||
Best practices for labelling
Be descriptive but concise
Keep labels clear and specific, but not verbose:
| ✓ Good | ✗ Avoid |
|---|---|
| “Age Group (years)” | “The age of the participant in years” |
| “Systolic Blood Pressure (mmHg)” | “BP_sys” |
| “Smoking Status” | “smoking_status_yes” |
Include units where relevant
Always specify units in parentheses:
var_labels <- list(
wt = "Weight (kg)",
ht = "Height (cm)",
bp_sys = "Systolic Blood Pressure (mmHg)"
)Use consistent formatting
Apply consistent capitalisation and punctuation across all labels:
# consistent approach to capitalisation
var_labels <- list(
ag_gp = "Age Group",
sm_status = "Smoking Status",
ed_level = "Education Level"
)Label factor levels clearly
Make factor level labels explicit and unambiguous:
df <-
data.frame(
education = sample(
x = 1:3,
size = 10,
replace = TRUE
) |>
factor(
labels = c(
"Primary school",
"Secondary school",
"University degree"
)
)
)Preserving labels in your workflow
R-native formats preserve labels
Labels are preserved when saving and reloading with .Rds or .RData:
CSV files lose labels
Labels are lost when reading from CSV or other text formats. To preserve labels, use one of these approaches:
Use R-native formats like
.Rdsor.RDataRe-apply labels after reading CSV files
Store labels separately in a data dictionary and apply them in your analysis script
See also
vignette("table_or")- formatting results tables and exporting with {gt}vignette("check_or")- diagnostics and model validationlabelled - comprehensive labelling package
haven - import labelled data from SPSS, Stata or SAS
Hmisc - labelling and statistical functions commonly used in epidemiology and biostatistics